Chai Shots #005: Miscalibrated Lipids
Why PCSK9 Inhibitor May Not Work for South Asians and Lp(a)
If you’ve been following lipid management guidelines, you’ve probably heard about PCSK9 inhibitors, the “miracle drugs” that cut LDL-C by 50-60%. Repatha, Praluent, and the newer siRNA agent inclisiran have transformed cardiovascular prevention.
So here’s the uncomfortable question to ask: What if this entire therapeutic strategy was optimized for the wrong population?
Lipids are heavily genetics driven, and this is something we’ll explore through research.
Two studies published in the past six months suggest exactly that, lets dive in.
Yes…Grab some chai.
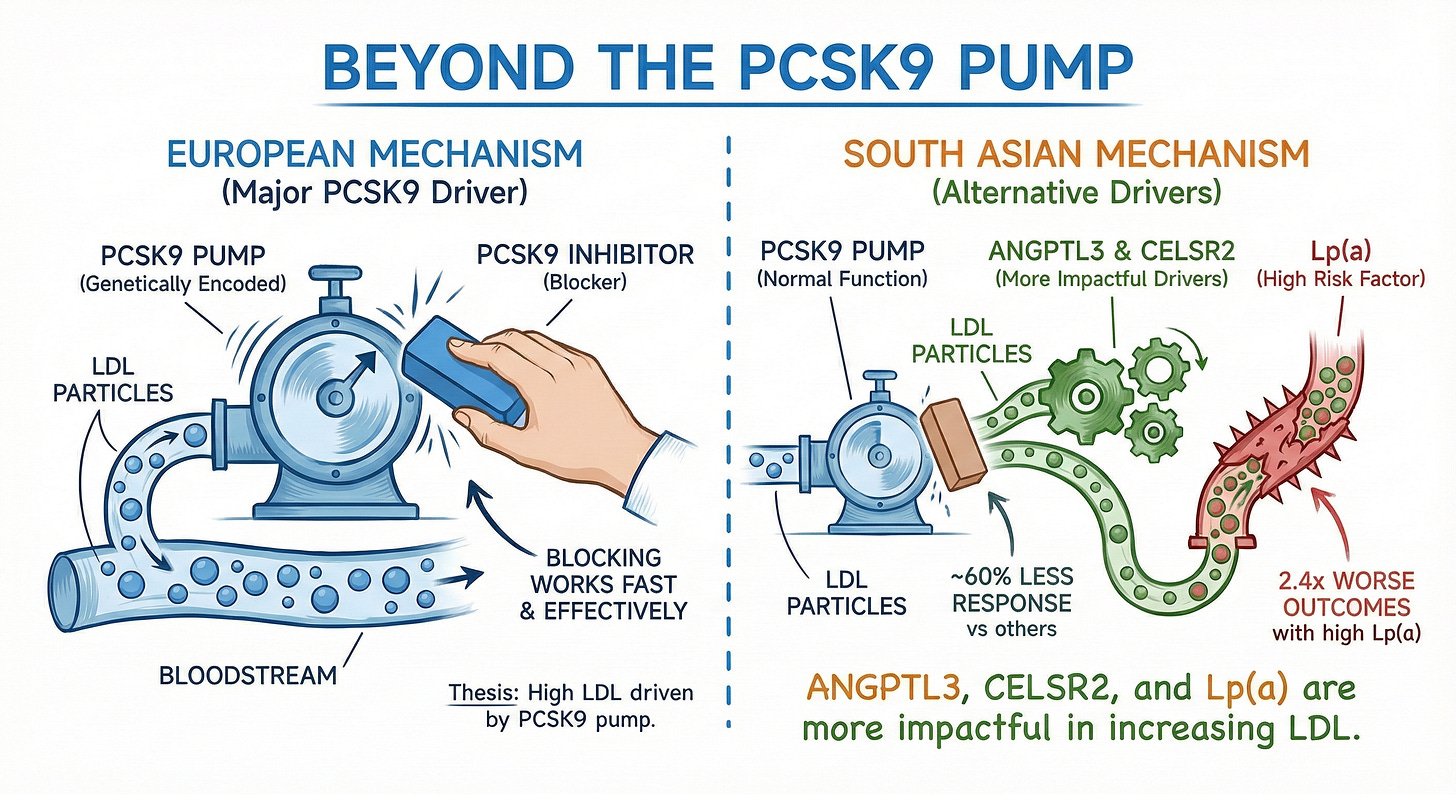
The PCSK9 Pump Problem
A Mendelian randomization study from The Lancet Regional Health – Southeast Asia examined the “proteomic architecture” of lipid metabolism across ancestries. The headline finding: PCSK9’s effect on LDL-C is 57% weaker in South Asians compared to Europeans.
The numbers are striking. The beta coefficient for PCSK9’s effect on LDL-C was 0.37 in Europeans versus just 0.16 in South Asians (95% CI: 0.11–0.21). This isn’t a subtle difference, it’s a fundamentally different relationship between the protein and the lipid.
The study identified why. In Europeans, PCSK9 functions like a master pump, genetically encoded variation in PCSK9 strongly predicts LDL-C levels and cardiovascular outcomes. Block the pump, solve the problem.
In South Asians, the pump is still there, but it’s not the main driver. Instead, three alternative pathways emerge as more impactful:
CELSR2 — This gene showed novel causal association with coronary artery disease specifically in South Asians, with strong Bayesian colocalisation evidence (PPH4 >80%) across multiple lipid fractions. (see table below)

ANGPTL3 — Its causal role in South Asian dyslipidemia was confirmed, with strong colocalisation with all lipid fractions except HDL-C.
Lp(a) — The genetic wildcard that keeps showing up in South Asian cardiovascular research, with confirmed colocalisation with LDL-C and total cholesterol.

The Genetics Deep Dive
This is where it gets interesting for those who’ve had pharmacogenomic testing or are considering it. One of my favorite areas to write about.
PCSK9 Variants: Population-Specific Effects
The R46L variant (rs11591147) is the poster child for PCSK9 loss-of-function genetics. According to research published in The New England Journal of Medicine, in Europeans:
Carriers have ~15% lower LDL-C
47-50% reduced risk of coronary heart disease
Enhanced response to statin therapy
Lower Lp(a), lower VLDL particle concentrations, lower inflammatory markers
The problem for South Asians: R46L is rare in Asian populations. Studies in Chinese populations found zero carriers of this protective variant. The allele frequency is approximately 3.2% in Europeans but <1% in South Asians, meaning this genetic “get out of jail free card” is largely absent in our population.
Other protective PCSK9 loss-of-function variants (Y142X, C679X) are similarly rare or absent outside European and African-American populations.
SOmething to think about. If you’re South Asian and your PCSK9 inhibitor isn’t hitting targets as expected, this may (stress on may) explain why. The genetic variants that predict excellent response are underrepresented in our population.
The 1p13 Locus: CELSR2/SORT1
The chromosome 1p13 locus is one of the strongest genetic signals for LDL-C and CAD in GWAS studies. The key variant is rs12740374, which:
Creates a C/EBP transcription factor binding site
Alters hepatic expression of SORT1 (sortilin-1), affecting VLDL secretion and LDL clearance
The minor (protective) allele increases SORT1 expression up to 10-fold
Here’s what makes this South Asian-relevant: The C4D Consortium found that the effect size of SORT1-related variants on CAD was significantly lower in South Asians than Europeans. What’s protective in Europeans may be neutral, or potentially harmful, in South Asians due to different linkage disequilibrium patterns and gene-environment interactions.
The Lancet SEA study specifically identified CELSR2 as having novel causal relevance in South Asians, with strong colocalisation across LDL-C, HDL-C, total cholesterol, and non-HDL-C. This suggests the entire hepatic lipid handling apparatus at this locus may function differently in our population.
Key SNPs to know at this locus:
rs12740374 (primary causal variant)
rs629301 (CAD risk haplotype)
rs599839, rs646776, rs660240 (all in strong LD)
LPA Gene Variants: The Lp(a) Story
Lp(a) genetics are notoriously complex, but here’s what matters clinically:
KIV-2 Repeat Number: The number of Kringle IV type-2 repeats in the LPA gene is inversely correlated with Lp(a) levels. Fewer repeats = smaller apolipoprotein(a) isoform = higher Lp(a) = higher risk. This is ~90% heritable and determines your lifetime Lp(a) trajectory.
rs10455872 (intronic SNP) and rs3798220 (causes amino acid change I4399M in the protease domain): These are the two most commonly tested Lp(a)-associated variants. According to the Journal of the American Heart Association, in Europeans, they:
Tag small apo(a) isoforms
Predict elevated Lp(a)
Associate with 1.47x and 1.68x increased MI risk, respectively
Combined genotype score shows 4.87x risk with two or more variant alleles
But here’s an interesting point for SAs (and something that illustrates precision therapy eloquently)
rs3798220 does NOT predict elevated Lp(a) in South or East Asians. A study published in Atherosclerosis examining this variant across ethnicities found:
The variant allele frequency in Indian CAD cases was only 0.15%
In Asians carrying rs3798220, there was no association with small KIV-2 alleles or elevated Lp(a)
The haplotype structure is different, the same SNP sits on a different genetic background
This has major implications: Commercial genetic tests using rs3798220 and rs10455872 to predict Lp(a) risk will generate high rates of false negatives in South Asian populations.
For rs10455872 specifically: According to recent comprehensive reviews, the minor allele is absent or extremely rare in East Asian and South Asian populations. If you’re South Asian and tested “negative” for Lp(a) risk variants, you still need to actually measure your Lp(a) level.
What to do: Get your Lp(a) level measured directly (in nmol/L, not mg/dL for accuracy). Don’t rely solely on genetic proxies developed in European populations.
ANGPTL3: The Emerging Target
ANGPTL3 inhibits lipoprotein lipase and endothelial lipase, increasing circulating triglycerides and LDL-C. Loss-of-function mutations cause familial combined hypolipidemia, remarkably low levels of all lipoproteins with no apparent pathology.
Key SNPs:
rs6690733 — The C allele increases ANGPTL3 expression and is associated with higher TC and LDL-C and increased ischemic stroke risk
rs12563308 — The C allele is protective, associated with lower TC and LDL-C
The Lancet SEA study confirmed ANGPTL3’s causal role in South Asian dyslipidemia with strong genetic evidence. This is significant because ANGPTL3 inhibition works regardless of LDL receptor function, it’s a fundamentally different mechanism than statins or PCSK9 inhibitors.
Evinacumab (Evkeeza) is already FDA-approved for homozygous FH. Given the weaker PCSK9 effect in South Asians, ANGPTL3-targeted therapies may prove particularly relevant for our population.
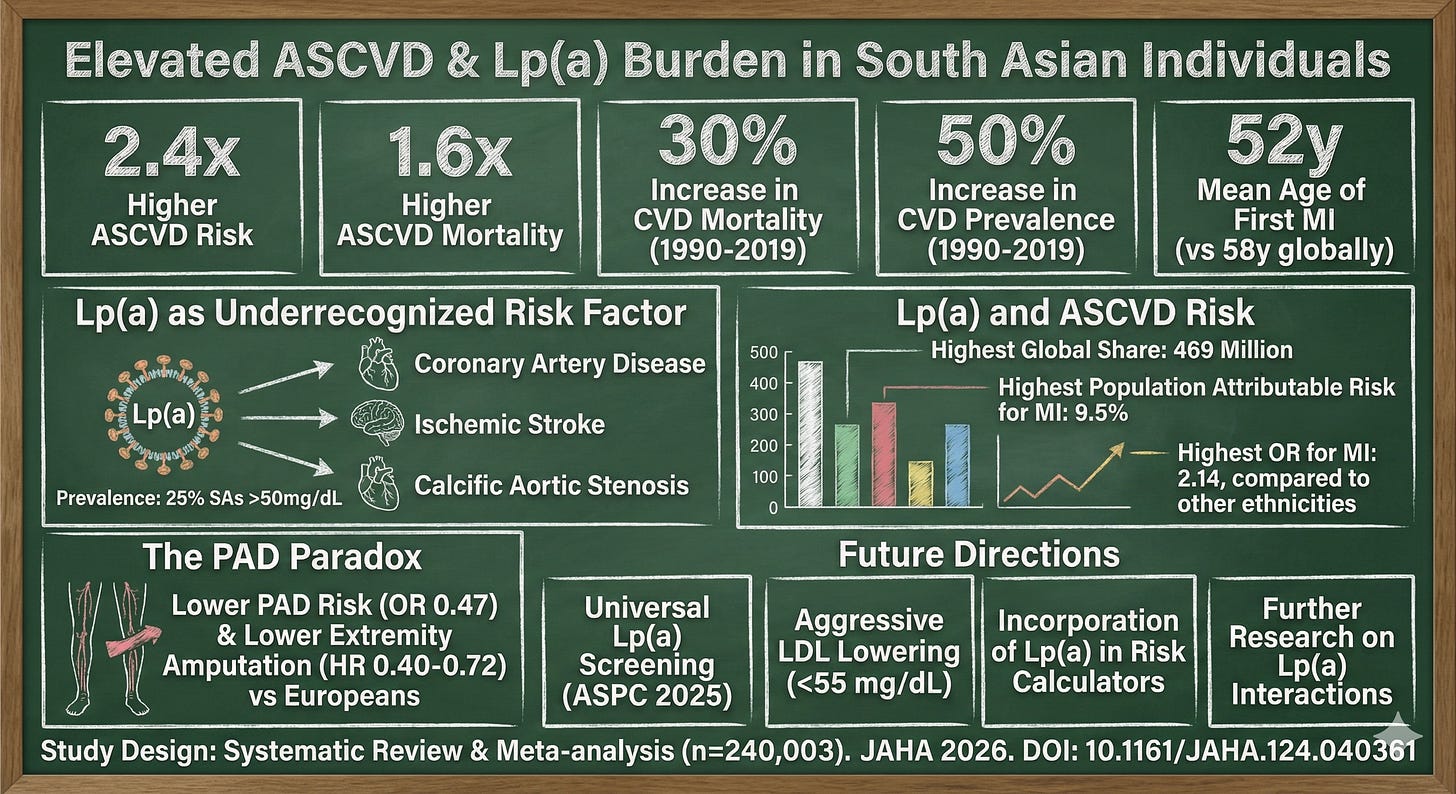
Lp(a): Same Test, 2x the Risk
The INTERHEART study and subsequent analyses published in the Journal of the American Heart Association confirmed what cardiologists have suspected: elevated Lp(a) confers substantially higher MI risk in South Asians compared to Europeans.
Same biomarker. Same laboratory cutoff. Completely different clinical meaning.
Lp(a) matters. Siginificantly.
A thirty-year study published in JAMA this week showed very high Lp(a) levels were associated with 30-year risk of cardiovascular disease among healthy women.
According to the National Lipid Association, the prevalence of elevated Lp(a) (>50 mg/dL) is higher in South Asians (~25%) versus Europeans (~20%), but prevalence alone doesn’t explain the risk differential. There’s something about how Lp(a) interacts with South Asian vascular biology that amplifies its atherogenic potential, possibly related to our tendency toward smaller coronary vessels, higher inflammatory burden, and different lipoprotein composition.
2026 is be a big year
Especially for treatments that matter for South Asians
The next 12-18 months will be transformative for Lp(a). According to the American College of Cardiology and recent updates in Pharmacy Times:
Pelacarsen (antisense oligonucleotide) — The phase 3 Lp(a)HORIZON trial has enrolled 8,325 patients with established CVD. Lowers Lp(a) by ~80%. Results expected early 2026.
Olpasiran (siRNA) — Phase 3 OCEAN(a) trial (NCT05581303) has completed recruitment of over 7,200 patients. Phase 2 showed 94% Lp(a) reduction. Results expected December 2026.
Lepodisiran — Phase 2 ALPACA data showed >90% Lp(a) reduction with a single 400 mg dose, persisting for over 360 days. Phase 3 ACCLAIM-Lp(a) trial currently enrolling 12,500 participants.
Muvalaplin — Oral Lp(a) inhibitor showing 86% reduction in phase 2. Yes, oral.
The critical question.
Will these drugs show the same efficacy across ancestries? The genetic architecture differs significantly. Early data suggest efficacy isn’t dramatically affected by baseline isoform size or common LPA SNPs, encouraging, but real-world South Asian representation in these trials matters.
What This Means for Treatment
If you’re South Asian with dyslipidemia, the standard algorithm may be insufficient.
The precision approach:
Get Lp(a) tested directly. Not negotiable. Don’t rely on genetic proxies, the SNPs don’t work in our population. It’s a one-time test (levels are genetically fixed), and if elevated, it fundamentally changes your risk calculus.
Push harder on ApoB. ApoB captures all atherogenic particles, LDL, VLDL remnants, and Lp(a). Given our alternative drivers, ApoB may be a more meaningful target than LDL-C alone. Consider a target <60 mg/dL if high risk, <80 mg/dL for primary prevention.
Don’t abandon PCSK9 inhibitors, but manage expectations. 57% weaker doesn’t mean zero effect. These drugs still work; they just may not be the home run they are in Europeans.
Watch the ANGPTL3 space. Given the confirmed causal role in South Asian dyslipidemia and the PCSK9-independent mechanism, this pathway may prove particularly relevant.
Consider advanced lipid testing. LDL particle number, ApoB, Lp(a), oxidized phospholipids, these give a more complete picture than standard lipid panels.
This is what precision medicine actually looks like. Not personalized wellness marketing, but hard data showing that ancestry-agnostic guidelines leave specific populations undertreated.
The proteomic architecture of lipid metabolism isn’t uniform across humanity. We evolved under different dietary pressures, different infectious disease burdens, different metabolic demands. The genetic variants that protect Europeans from heart disease are often rare or absent in South Asians. The SNPs used to predict drug response don’t work in our population. The effect sizes of therapeutic targets differ by 50% or more. Yet we still barely have SNPs and other omic structures that can be harnessed for south asians, especially the diaspora underrepresented in populations.
The playbook is being rewritten. We’re the authors now. Make sure you’re reading the right edition.
-Omar
This is educational content. Work with your physician on any medication decisions. If you found this useful, please share it with a family member who should know their Lp(a) status or would be interested in learning more about their body. We are also working on building something for the South Asian diaspora. More on this next week!